So, welcome back to my intermittent live-blog of my adventures in forming a non-profit research institute in order to function as an independent scholar. I’ve written a couple of times before: about my own goals for the enterprise, and about the things that an independent scholar will most be in need of.
One of the things, of course, that an independent scholar needs is colleagues. Depending on the nature of your research, you might be able to do the day-to-day work (math and programming, in my case) entirely on your own, but unless you are a very special sort of misanthropic genius, you need interaction with a set of colleagues. Sometimes you will want to take on collaborative projects that require the expertise of more than one person, but even more, you need knowledgeable people to bounce ideas off of, people who will ask the critical questions that make your work better, or who will drop some jewel of knowledge that lets you see the problem you’ve been working on in an entirely new way.
Now, in principle, much of this can be accomplished on the internet, but I am wondering if there are not certain types of information that more or less require face-to-face contact.
Last week, I was at a “catalysis meeting” at NESCent (the National Evolutionary Synthesis Center) on genomic imprinting. The meeting was superb. It had excellent people who work on the problem from all different perspectives: theorists and experimentalists, molecular and developmental biologists, mouse people, marsupial people, bee people. I learned a ton, and, perhaps more importantly, I learned of the existence of a bunch of things that I didn’t know. I still don’t know them, but now I know that I should know, and I know where to start looking, and whom to ask for help when I get stuck.
 |
| As an aside, I also had the chance to meet Craig McClain, Assistant Director of Science at NESCent and doyen of the group blog Deep Sea News. He was as nice as their blog is awesome.
Some people say that biologists grow to resemble the organisms that they study. You be the judge. |
You might think that meetings like this are particularly efficient for transmitting information, but that you can accomplish the same thing through more aggressive and far-reaching readings of the literature. After all, the organizers of the meeting were able to find these people. In principle, I could just get all of their papers and read them carefully, referring to textbooks on biochemistry or mammalian physiology whenever there was something I didn’t understand.
But I’m not sure that would actually work.
The thing is, some of the most important pieces of information I got at the meeting were things that are not written in papers, or perhaps anywhere, nor are they likely to be. For example, there were a number of people there who have spent years working with lab mice. They have observed thousands and thousands of crosses (e.g., the outcome of a mother of one mouse strain mating with a father of a different mouse strain). This has given them a deep knowledge of what does and does not happen in these crosses, as well as a sense of how sensitive different traits are to the details of the experimental procedure.
An interesting thing was that there were certain results from the scientific literature that none of these people believe, because they are not consistent with their own observations. Now, no one has gone and written a rebuttal letter, or published a set of negative results contradicting the original papers. They have all just sort of implicitly agreed that results using a certain technique, or sometimes results coming from a certain lab, are unreliable, and they move forward with their research as if those results did not exist.
So, there is this substratum of knowledge that is widespread among experts, but which does not find its way into print. In part, this is due to the thanklessness of writing response letters and publishing negative results. In part, I think, it results from a sense of decorum / political consideration. It is common for scientists to have opinions that whole swaths of research are garbage, and it is common for them to share this knowledge in conversation, particularly over beer. However, most are too cautious to put their genuine opinions down in writing — even in e-mail.
 |
| As the good folks at Gawker say, “Today’s gossip is tomorrow’s news!” |
Fundamentally, I don’t think that there is anything wrong with this arrangement, as it maintains a pretty high bar for calling someone out for doing bad science, but permits people to move forward with what they collectively perceive to be the best possible information. However, it does point to the importance of getting out there and interacting with people face to face. Otherwise, you may find yourself developing a whole research project that is predicated on some results that no one thinks are true.
I should note that this problem is not unique to the independent scholar. If you are working in a typical university department, there may not be anyone else in your department — or only a small number of people — whose research is close enough to your own that you share the same scuttlebutt. That is, no matter who you are, you need to make sure that you pursue opportunities to talk informally — and in person — with the people who care about the same things that you do.
One last observation from the NESCent meeting. This was the first scientific meeting I have attended under my official affiliation with the Ronin Institute. This meant that people would look at my name tag and ask me about it. I would tell them briefly about the idea and my plans for Ronin, and they were all very enthusiastic. The people who had come over from England, in particular, tended to comment on how very brave I was. After I got back, I came a cross this translation guide:
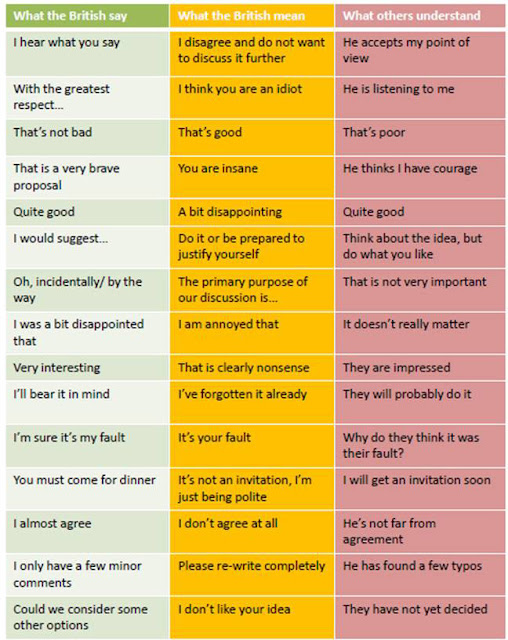 |
| If you work with anyone British, you should print this out and carry it around with you. It serves as a handy guide as to whether you need to be punching them in the nose. |
I’m going to assume that this is just wrong. Let’s posit that a better translation for “That is a very brave proposal” would be “Wow! You are a singular genius and an inspiration to children around the world! Also very sexy! Mee-yow!”










